PLOD1-kEDS Cure Roadmap
As Perlara quietly reemerged in the summer of 2021 as the first decentralized biotech cure consultancy, a family from Mexico City reached out to us asking for help.
Perlara 2.0 emerged last summer after a two-year hibernation. My experience with SDSUK and Julia Hawkins convinced me that Perlara could provide a valuable service to entrepreneurial cure-focused families and foundations. The first family to fill out a Guided Cures form wrote the following message:
”We are a family from Mexico City and my niece is diagnosed with Kyphoscoliotic Ehlers Danlos (PLOD1 gene), a rare and problematic genetic disease with less than 100 documented cases in the world.
Today, this disease has no cure and very limited research around it. It is very difficult for us to think that there may be an existing drug out there already that could help her, but due to limited research and funds it is not being looked into.
We are willing to do everything necessary to find a cure. Do you have some time available soon to schedule a call to discuss our opportunities? Please let me know if you need any further information on my niece and her detailed medical history, we would be more than happy to share it.”
PLOD1-Related Kyphoscoliotic Ehlers-Danlos Syndrome Cure Roadmap
Prepared for Health and Life Forever LLC by Perlara PBC
November 2021
VISION
Toward a future where people with PLOD1-Related Kyphoscoliotic Ehlers-Danlos Syndrome live healthy, fulfilled and complete lives. This multi-year, multi-modality drug development plan makes the scientific and commercial case for PLOD1-kEDS. It calls for rigorous testing of two therapeutic approaches and a plan for clinical trial readiness. This Roadmap envisions the following milestones and timelines:
In 4-6 months, the preliminary experimentation will be performed to assess the likelihood for success of the proposed therapeutic modalities. Drug repurposing and antisense oligo (ASO)-based precision medicine modalities will be tested, putting our theoretical assumptions about which treatment avenues offer the most long-term promise to the test.
In 7-12 months, we will have entered the validation stage of the promising therapeutic modalities. In-depth characterization of ASO and repurposed drug candidates will be underway. If the ASO-based precision medicine modality fails as a therapeutic avenue in the first 4-6 months, we will pivot to developing an enzyme-replacement (ERT) or gene therapy-based precision medicine strategy at this time.
In 16-24 months, we will have preclinical data packages for promising therapies in-hand so that we can pursue fundraising and partnership opportunities with biotech and pharmaceutical companies. We will also have a pre-IND meeting with the FDA to vet drug development plans. It is at this stage that we will expand the pioneer “n-of-1” program to conduct pivotal clinical trials to serve the community of PLOD1-kEDS patients writ large.
AUTHORS
Victoria M. Blake, PhD (Cure Guide, Perlara PBC)
Shiri Zakin, PhD (Cure Guide, Perlara PBC)
Ethan O. Perlstein, PhD (CEO of Perlara PBC & Maggie’s Pearl LLC)
REVIEWERS
We invite post-publication review of this Cure Roadmap here on our Substack.
EXECUTIVE SUMMARY
The PLOD1-kEDS Cure Roadmap marries the best of two complementary models of patient-driven drug development. Firstly, a mutation-agnostic approach that aims to identify therapies that bypass the gene’s function; and, secondly, a mutation-targeted approach that aims to restore protein function in key tissues. A blended strategy is necessary because of the inability to predict the success of any one therapeutic strategy for treating PLOD1-kEDS, especially given the involvement of multiple organ systems.
XXXXX’s family and the PLOD1-kEDS community are poised for capital-efficient, modality-diversified drug development. A PLOD1-kEDS knowledge base has been forged by years of disease biology insights. A multi-species array of cell and animal PLOD1-kEDS disease models already exist or can be quickly generated on a variant-by-variant basis. Straightforward detection of clinical endpoints allows for the generation of a standardized clinical trial protocol that will be vetted by regulatory agencies and pressure-tested in single-patient “n-of-1” pioneer studies.
One drug target -- LH1 -- has been robustly and reproducibly validated by both human and model organism genetics (Pinnell et al., 1972; Krane et al., 1972; Yeowell & Walker, 2000; Malfait 2018; Scietti et al., 2019; Takaluoma et al., 2007). Although the Ehlers-Danlos community has spent a great deal of resources on improving diagnostics, no large-scale drug discovery efforts are underway to target molecules like LH1 for treatment of PLOD1-kEDS.
This roadmap initiative was kicked off by Moises Askenazi, Emilio Gamus, Ethan Perlstein, Shiri Zakin, and Victoria Blake in September 2021. During the autumn months, we invited reviewers from academia, medicine, and industry to participate and have their voices heard. This Roadmap will continue to be transparently and rigorously reviewed “post-publication” throughout the winter months as we begin experimentation on potential therapeutic avenues.
The goal of the PLOD1-kEDS Cure Roadmap is to stimulate development of medicines that will treat (and potentially cure) PLOD1-kEDS. Perlara’s agile Cure Guide operating structure and a Slack-first, globally distributed team will enable research capacity building and preclinical proof-of-concept studies across multiple programs that will ultimately coalesce into a biotech startup ecosystem urgently developing new curative medicines.
The PLOD1-kEDS Cure Roadmap project will deliver:
Exploration of mutation-agnostic and gene-specific therapeutic modalities as potential treatments of the syndrome. These modalities include:
An unbiased drug repurposing screen using patient-derived fibroblasts.
Precision medicine treatments for restoring function of the LH1 enzyme that include:
An ASO-based exon-skipping therapy to mask the mutation-harboring portion of the gene and restore protein functionality.
A gene therapy delivered by non-integrating virus in somatic (not germline) cells to correct the mutation in the PLOD1 gene.
An enzyme-replacement therapy to deliver functional LH1 to key organs.
Outlining of a standardized clinical trial protocol that will be validated by regulators in order to lower the barrier to investment into specific therapeutic tracks.
Disease models, prognostic tests, clinical trial protocols and disease-modifying therapies developed for PLOD1-kEDS may have broader applicability and impact across many Ehlers-Danlos subtypes, other monogenic diseases, and may ultimately impact healthspan extension. For example, insights or discoveries leading to the rejuvenation or remodeling of extracellular matrix (ECM) in the context of diseases of aging in adults. This broader applicability will make investments into PLOD1-kEDS compelling beyond the stakeholders who are directly affected by the disease.
INTRODUCTION
What is PLOD1-related Kyphoscoliotic Ehlers-Danlos Syndrome?
XXXXX is a three year-old girl living in Mexico City living with PLOD1-related Kyphoscoliotic Ehlers-Danlos syndrome, which is a hereditary connective tissue disorder characterized by defects in collagen cross-linking. The prevalence of Ehlers-Danlos syndrome is between 1 in 2500 and 1 in 5000 births (Ghali and Burrows, 2019), though it is likely underdiagnosed given its phenotypic heterogeneity. In contrast, the prevalence of PLOD1-kEDS is 1 in 100,000 births (Rohrbach et al., 2011). To put these numbers in context, the threshold for a rare genetic disease is 1 in 2000 births, and the prevalence of 1 in 50,000 to 1 in 100,000 births approaches ultra-rare genetic diseases. Furthermore, though there are approximately 50 gene variants described as causative in PLOD1-kEDS (Scietti et al., 2019), XXXXX’s specific pathogenic variant is yet unreported in the scientific literature.
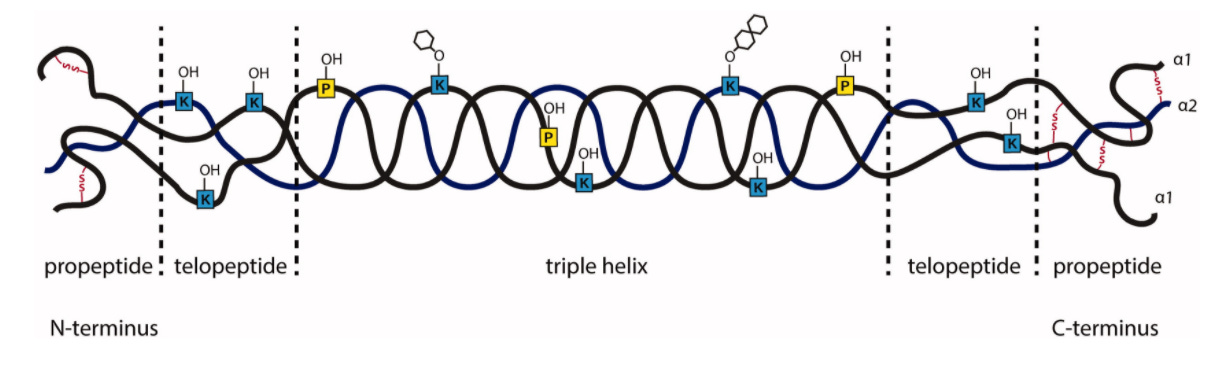
Kyphoscoliotic Ehlers-Danlos syndrome is a disorder caused by an autosomal recessive loss-of-function mutation in the PLOD1 gene, which encodes for Lysyl Hydroxylase 1 (LH1) enzyme. Lysyl hydroxylases, of which there are three isozymes, are responsible for hydroxylating specific lysines on the collagen fiber during collagen maturation. These hydroxylated lysines are sites of glycosylation and subsequent cross-linking of the collagen fiber, which controls the tensile strength of the extracellular matrix. Failure to properly cross-link the collagen fibers through this lysine hydroxylation pathway thus results in fragile tissues that make up organs throughout the body (Yeowell and Walker, 2000).
XXXXX's clinical presentation of PLOD1-kEDS demonstrates the drastic effect of LH1 deficiency on multiple organ systems. The most common features of the disorder are hyperextensible, fragile skin that bruises easily, joint hypermobility, severe hypotonia, progressive kyphoscoliosis, fragility of the sclera, and arterial fragility and risk for rupture. XXXXX exhibits all of these symptoms and notably experienced a right brachial plexus injury and an intraventricular hemorrhage in her right temporal lobe at birth. She has since had surgery to correct her injury and is being seen by a cardiologist, neurosurgeon, and opthamologist to monitor her disease progression. XXXXX is also in regular consultation with occupational and physical therapists to manage her hypotonia, joint hyperextensibility, and kyphoscoliosis.
The three LH isozymes are differentially expressed throughout the body and are not interchangeable, a physiological observation that is key in tailoring therapeutic strategies for treatment of PLOD1-kEDS. Notably, loss of LH2 results in Bruck Syndrome, another collagen cross-linking disorder, whose clinical presentation presents as osteogenesis imperfecta and joint contractures rather than the severe hypotonia and joint hyperextensibility seen in PLOD1-kEDS. Though both conditions affect several organ systems, the differences in clinical presentation stress the importance of the differential expression of the PLOD genes (Uzawa et al., 1999) in the development of therapeutic modalities.
Diversified portfolio of therapeutic modalities
The inability to predict the success of any one treatment modality necessitates a diversified portfolio of mutation-agnostic and mutation-targeted strategies that are embodied in distinct but combinable therapeutic modalities.
We are fortunate in that cell-based and animal models are readily available for testing any one of the therapeutic avenues proposed. With these models in hand, the PLOD1-kEDS Cure Roadmap contemplates advancing toward new medicines along two main therapeutic tracks: (1) unbiased drug repurposing and (2) nucleic acid therapies. If the early prognostic markers of success for the nucleic acid strategy are unfavorable, we will then consider gene replacement and enzyme-replacement therapies as alternative mutation-targeted treatment avenues.
Ultimately, we predict that XXXXX will require some form of combination therapy consisting of a mutation-agnostic medicine, e.g., a repurposed drug that bypasses the PLOD1 gene function, and a mutation-targeted medicine, e.g., an exon-skipping antisense-oligo (ASO) therapy that restores LH1 function. In a combination therapy scenario, an orally bioavailable small molecule medicine that reaches all cells of the body may be combined with another therapeutic modality that has restricted tissue tropism or restricted tissue delivery.
Other therapeutic modalities need de-risking
Enzyme-replacement and gene therapy approaches need de-risking and therefore are not considered in Year One activities. These modalities will be revisited contingent on the outcome of drug repurposing and antisense oligonucleotide projects.
Balancing drug-repurposing and precision-medicine approaches
There are many challenges with treating PLOD1-kEDS that necessitate a multi-modality treatment approach. The LH1 enzyme performs a critical role at the end of a collagen maturation pathway, and there are no known downstream effectors that can be upregulated to compensate for the failure to hydroxylate lysines on collagen. Although other LH enzymes are thought to compensate for loss of LH1 (Eyre et al., 2002), these enzymes are differentially expressed, and misexpression of these enzymes is associated with many cancers (Qi and Xu, 2018). Finally, we cannot predict the deliverability challenges of a novel therapy, especially given that the affected tissues range from the skin to smooth muscle vasculature as well as poorly-vascularized microenvironments in some cartilaginous tissues.
THERAPEUTICS READINESS
The development of new medicines requires disease models, patient-derived biosamples, and a deep mechanistic understanding of pathophysiology that identifies disease modifying drug targets, including the monogenic driver gene itself. The primary function of LH1 protein encoded by the PLOD1 gene is in a fundamental biological process called lysyl hydroxylation, a necessary step in collagen crosslinking leading to healthy extracellular matrix or ECM. Accordingly, simple cellular models (patient-derived cells and other tissue-specific cell lines) can demonstrate very well the degree to which a given intervention normalizes, for example, a cellular phenotype such as collagen maturation. Higher animal models then show how failure to hydroxylate lysines on the collagen fiber (either in quantity or quality) emerges as a phenotype, which varies between species and between tissues within a species.
But how much does the particular manifestation in, say, mouse vasculature, tell us about humans? Divergence of pathophysiology between even closely related species can occur in spite of overwhelming genome sequence conservation. As a result, all information gained from disease models must be multi-factor authenticated and, as such, consensus of mechanism and functional recovery in multiple models is needed to provide the best chance of therapeutic benefit.
Disease models
Cell-based and animal models of PLOD1-kEDS allow for the experiments necessary to identify potential therapies for PLOD1-related kEDS. We are lucky in that these models have already been developed and thoroughly characterized through basic research. Furthermore, relevant cell lines (that are not patient-derived) and animal models are available to the scientific community for further experimentation and in our case, for developing therapeutic strategies for PLOD1-kEDS patients like XXXXX.
Below is a description of the relevant PLOD1-kEDS disease models.
Cell-based models
Patient-derived fibroblasts are a cheap and efficient in vitro experimental system in which to test both mutation-agnostic and mutation-targeted PLOD1-kEDS therapies. These cells are inexpensive to maintain, easy to culture, and are amenable to the high-throughput assay development needed for drug repurposing screens. Fortuitously, fibroblasts are derived from an organ rich in type I collagen which is affected by PLOD1-kEDS, so we know that the effect of any given therapy measured in these cells is applicable for treating the tissue from which they are derived.
Another cell type we can employ during the validation stage of PLOD1-kEDS therapies are vascular smooth muscle cells (VSMCs) derived from aorta. These are commercially available from Life-line Cell Technology, CA. We are especially interested in testing candidate therapies in this tissue because of the vascular fragility present as a result of PLOD1-kEDS. Though no PLOD1 knock-out VSMCs are available, one can use an siRNA to knock-down target PLOD1 mRNA and recapitulate the PLOD1-kEDS phenotype (Koenig et al., 2021).
A humanized PLOD1 knock-out mouse model
PLOD1 gene knock-out in mice recapitulates the PLOD1-kEDS human disease phenotype (Takaluoma et al., 2007). The mice are flaccid, have gait abnormalities, and display abnormal collagen fibril morphology in the aorta and skin. 15% of the mice died due to aortic rupture and all mice studied showed signs of degeneration in vascular smooth muscle cells. Hydroxylysylpyridinoline cross-links were decreased in all tissues studied.
We propose utilizing this model system if an organismal validation checkpoint for promising repurposed drugs or precision medicine modalities is needed. The humanized PLOD1-/- mice were generated by the Myllyharju lab in 2007 and are available through Intrafrontier.
Additional model organisms available for PLOD1 study
PLOD1 is orthologous to the CG6199 gene in Drosophila melanogaster. Studies on this gene demonstrate that disruption of the ninth exon with a P element insertion causes embryonic lethality (Graur et al., 2008). Importantly, several alleles that disrupt CG6199 generated by the Berkeley Drosophila Genome Project are publicly available for further study (Bellen et al., 2004).
Although there is only one gene that encodes a lysyl hydroxylase in C. elegans, it is highly similar to the vertebrate LH proteins and contains the same essential motifs needed for enzyme activity (Norman and Moerman, 2000).
Clinical Trial Readiness
Measurement of urinary pyridinolines is a widely-used assay that detects degraded, but still cross-linked, collagen which is routinely used in diagnosing PLOD1-kEDS (Yeowell and Steinmann, 2018). The ratio of hydroxylysyl pyridinoline (HP) to lysyl pyridinoline (LP) in urine is typically 3:1 in healthy individuals but is reversed to a 1:5 ratio in PLOD1-kEDS patients (Eyre et al., 2002).
Clinical trialists will monitor dosing and efficacy of a repurposed drug or promising ASO by continuously monitoring the ratio of HP:LP in the urine of PLOD1-kEDS patients. Because urinary pyridinolines are degradation products from low turnover tissues (such as bone), renal stem cells will be obtained from urine and cultured (Zhou et al., 2012) in order to directly and non-invasively assess changes to collagen crosslinking in a high turnover tissue. Here, we will take advantage of the observation that under-hydroxylated collagen exhibits faster electrophoretic migration, we probe lysyl hydroxylation in a high-turnover tissue by running alpha-collagen isolated from renal stem cells on an SDS-PAGE (Jarisch et al., 1998).
As a final assessment of drug efficacy, clinicians will also probe gingival bleeding In order to indirectly assess the tensile strength of a high-turnover cartilaginous tissue. PLOD1-kEDS patients, as well as many other individuals with EDS, exhibit vascular fragility and bruise easily. Measurement of gingival bleeding (Newbrun, 1996) is therefore a straightforward and clear indication of changes to tensile strength of the vasculature in PLOD1-kEDS patients.
The clinical endpoints enumerated in this section will be modified throughout the Roadmap project. Specifically, the data generated during Year One activities will serve as the basis of a pre-Investigational New Drug (IND) meeting request submission. We will receive responses from FDA on what exact safety measures and clinical assessments will be needed for a PLOD1-kEDS experimental therapy.
Standardized Clinical Trial Protocol
The experiments performed during Year One of this Roadmap will build a foundation for a pre-Investigational New Drug (IND) meeting request for the treatment of PLOD1-kEDS. Feedback from the pre-IND meeting will give us clarity on what kind of clinical study will best serve patients in the development of therapies.
Therapeutic Tracks
THERAPEUTIC TRACK I: PRECISION MEDICINE
Precision medicine is defined as a therapy that uses a patient’s specific genetic or molecular profile to prevent, diagnose, or treat a disease (National Cancer Institute). In this therapeutic track, we explore strategies that span different timelines. The first therapeutic approach we will explore is a variant-specific targeting strategy whose efficacy we can evaluate within the first year of our research program. The other therapies we propose will serve as a back-up long-term plan for treating PLOD1-kEDS in case our first precision medicine strategy fails.
Exon-Skipping
Exon skipping is an antisense oligonucleotide or ASO-based treatment whereby the exon harboring the disease-causing mutation is omitted from the mature mRNA, therefore restoring translation of a functional protein (Li et al., 2018). Specifically, the ASO targets donor or acceptor splice sites in the affected exon on the pre-mRNA and thereby changes the splicing pattern of the mature mRNA. Exon-skipping therapies are FDA approved and used for treating Duchenne muscular dystrophy (Duchenne MD) and spinal muscular atrophy (SMA).
ASO-mediated exon skipping is a very attractive therapeutic avenue for treating XXXXX’s PLOD1-kEDS because of her specific variant of the PLOD1 gene. She has a guanine deletion in exon 5, resulting in a frameshift and early termination of the peptide. The PLOD1 gene has 19 exons and the encoded LH1 is 729 amino acids, where the lysyl hydroxylase domain maps to the C-terminus of the protein (Scietti et al., 2019). Though many of the documented PLOD1 variants result in an early termination of the peptide sequence, many mutations map specifically to the lysyl hydroxyl domain (Heikkinen et al., 1997, Yeowell et al., 2000). Importantly, the SiMPLOD database of collagen lysyl hydroxylase (LH/PLOD) enzyme variants does not have any record of a deletion in exon 5 of PLOD1 causing PLOD1-kEDS or other LH-related disease (Scietti et al., 2019). According to this database and the inferred functional domains based on the solved structure of LH3 (Scietti et al., 2018), exon 5 of PLOD1 maps to the glycosyltransferase (GT) domain of LH1, though no reports of in vivo GT activity have been published for this enzyme. However, there is one report of LH1 glucosylgalactosyltransferase activity in vitro which warrants further investigation (Koenig et al., 2021).
We propose testing the exon-skipping therapeutic strategy in patient-derived fibroblasts. Before testing ASO efficacy, we want to determine if a PLOD1 mRNA missing exon 5 will yield a functional LH1 protein. To that end, we will test an allelic series of PLOD1 sequences that lack exons that do not map to the lysyl hydroxylase domain of the protein. The rationale behind the allelic series is that if a protein lacking only exon 5 is non-functional, perhaps a protein lacking more than just the fifth exon folds correctly and restores lysyl hydroxylase function. Having a comprehensive understanding of which protein variants are functional is significant for two reasons. One, if we have multiple functional shortened versions of LH1, this maximizes the potential for designing an effective exon-skipping ASO by virtue of having more splice sites to test. Secondly, these data represent a basis for treatment of PLOD1-kEDS variants where mutations are not confined to the fifth exon, therefore serving a much wider population of PLOD1-kEDS patients whose mutations cause early termination of the LH1 protein.
The allelic series are a straightforward set of go/no go experiments where the necessary transgenes will be ordered from a company like Twist Biosciences. We will assay for restored lysyl hydroxylation through immunostaining collagen to look for a restoration to fibril morphology (Micha et al., 2019). Given the observations that under-hydroxylated collagen exhibits faster electrophoretic migration, we can further probe lysyl hydroxylation by running alpha-collagen on an SDS-PAGE (Jarisch et al., 1998).
If any exon-skipped proteins from the allelic series are functional in restoring lysyl hydroxylation, we will proceed with designing ASOs against splice donor and/or acceptor sites in exons we want to exclude from the final protein sequence. We will utilize eSkip-Finder, a machine learning-based tool for identifying ASO sequences for optimal exon-skipping (Chiba et al., 2021). We will choose 2-3 oligos per target exon designed by this program to then test in fibroblasts, where we will use a Western blot assay for LH1 to detect restored protein synthesis in treated cells. If we identify an ASO (or several) that bypass the frameshift mutation in exon 5, we will utilize the assays used in the allelic series experiments to determine if restoration of collagen cross-linking has taken place.
The exon-skipping therapeutic modality is attractive for two key reasons. One, this therapy does not affect the expression pattern of the PLOD1 gene. This means that, assuming that there are no issues with delivery of the ASO, functional LH1 will only be produced in the tissues where PLOD1 is expressed and not in organs where the gene is off. Preserving the gene’s regulation is critical in avoiding mis-expression that may lead to cancer (Qi and Xu, 2018). The other advantage of this modality is the efficiency with which we can compile a data package to validate or reject the therapeutic strategy. Data from the in vitro experiments described above are sufficient to convince a potential biotech partner that this modality is a viable option for commercial development and clinical trials. This is in contrast to the in vivo mouse experiments necessitated by a gene therapy avenue, or manufacturing costs and quality control needed for investigation of an enzyme replacement therapy. Conversely, if this treatment modality fails, we will have invested minimal time and research dollars to learn that an alternative precision medicine-based modality would better suit PLOD1-kEDS treatment.
If this proves to be a successful therapeutic avenue, we will explore what ASO backbone chemistries, base modifications, and/or even attachable ligands are needed to address drug delivery to target tissues beyond the cutaneous route of administration.
Precision medicines modalities to explore in the event that exon-skipping modality is not viable
As previously mentioned, we propose alternative avenues to explore in the event that the ASO-based exon skipping strategy fails. We have outlined what we see as reasonable avenues to pursue in the present day. However, research that enables the development of these modalities is moving at breakneck speed, and we will have to reassess the clinical landscape and therapeutic options if we decide to explore the following treatment modalities.
Enzyme-replacement Therapy
We propose exploring Enzyme-replacement Therapy (ERT) as a therapeutic avenue for promoting healthy collagen maturation by targeting recombinant LH1 protein to key tissues affected by PLOD1-kEDS. ERT is the current standard of care for lysosomal storage disorders (Kakkis, 2002 and Parenti, Andria, and Ballabio, 2015).
The LH1 enzyme can be expressed as a recombinant protein (Krol et al., 1996) which makes ERT an attractive therapeutic modality for the treatment of PLOD1-kEDS. The challenge we face in pursuing this therapeutic avenue is efficient delivery of the enzyme to the key tissues affected by the absence of LH1. We propose enzyme-targeting to the smooth muscle vasculature and skeletal muscle, since these are the most affected tissues with potentially life-threatening consequences for PLOD1-kEDS patients. Administering recombinant human α-glucosidase to infants with Pompe's disease (Winkel et al., 2003) ameliorated skeletal and smooth muscle defects, and has improved survival rates and clinical outcomes of individuals living with the disease. Although bioavailability and diffusion across the blood-brain barrier remains a core challenge for recombinant enzyme delivery (Prater et al., 2012), we see the success in treatment of Pompe’s disease as a proof-of-concept for PLOD1-kEDS.
Gene therapy
Of the proposed therapeutic strategies for PLOD1-kEDS, gene therapy is the most nascent in its development and will likely take the most time to get to a treatment in-hand. To explore this modality, we met with an expert at Amryt Pharma, which is developing a non-viral, topical gene therapy for the treatment of Epidermolysis bullosa (EB), another rare collagen disease. We discussed the potential for using a gene therapy like what Amryt is developing for EB to target tissues such as the skin, gut, and eye. Importantly, topical administration of a gene therapy to these tissues without using a non-viral delivery system is attractive because it is safer than its virus-based counterpart (Wahane et al., 2020). This connection we made with Amryt was significant in that it opened a door for a future partnership (regardless of therapeutic modality) once we have a preclinical data package and feedback from the FDA for a pre-investigational new drug in-hand.
Gene therapy is currently being used for treating genetic disorders of the eye (Bainbridge et al., 2006). Though the eye is not among the most drastically affected organs in the clinical presentation of PLOD1-kEDS, introducing a functional copy of LH1 in these tissues can result in a significant improvement in the quality of life for someone like XXXXX.
XXXXX’s PLOD1-kEDS puts her at risk for retinal detachment and globe rupture, and as a consequence needs to be extremely protective of her eyes. Given the success and FDA approval of LCA2 gene therapy (Askou et al., 2021), we propose repurposing this approach for delivering a functional copy of PLOD1 to the eye. Unlike ERT, once the functional copy of PLOD1 has been incorporated into a cell’s genome, that gene is then subject to regulatory processes that will ensure proper distribution of LH1 in the eye.
THERAPEUTIC TRACK II: DRUG REPURPOSING
Drug repurposing is defined as identifying a new use for an already approved drug, be it a century old drug or a drug that was just recently approved. Drug repositioning is defined as salvaging an experimental drug that cleared the initial hurdle of early-stage safety studies but failed in late-stage efficacy studies for its intended disease indication.
The Broad Repurposing Hub (commercially available as the SPECS library) is an ideal launchpad for identifying drug repurposing and drug repositioning candidates. All validated PLOD1-kEDS disease models will be screened against this collection as part of a coordinated multi-species high-throughput drug screening campaign.
Based on a review of the literature, it is likely that the other two LH isozymes may be partially compensating in part for the loss of LH1 in PLOD1-kEDS patients (Eyre et al., 2002). Given that the LH enzymes are Fe2+-, ascorbate-, and α-ketoglutarate-dependent (Ishikawa and Bächinger, 2013), it is possible that a drug that modulates the availability of any of these cofactors may alleviate the loss of LH1 by boosting the activity of a compensatory lysine hydroxylating enzyme in the affected tissues. There may also be yet-unidentified molecules in tissues lacking LH2 and LH3 that may compensate for loss of LH1 whose compensatory mechanisms may be revealed by an unbiased drug repurposing screen.
Patient-derived primary cell lines in unbiased image-based screens
The first step in pursuing a drug repurposing approach is a primary phenotypic screen on patient-derived skin fibroblasts. Skin fibroblasts are a well-established model for studying PLOD1-kEDS given that they are derived from an affected tissue and are an inexpensive model system for high-throughput studies. We also plan to follow up on leads from our drug repurposing screens in fibroblasts derived from XXXXX’s parents, who are heterozygous for the causative PLOD1 mutation. This will serve as a way to rule out drugs that may cause hyper-hydroxylation and subsequent aberrant collagen cross-linking in healthy tissues.
We propose the following workflow for our proposed screen in fibroblasts. The primary readout for drug efficacy would entail using a fluorescently-conjugated antibody to detect collagen fibrils and a separate antibody to detect hydroxylated lysine residues in the extracellular matrix. The anti-collagen antibody would detect for corrections in aberrant directionality of collagen fibril organization (Micha et al., 2019) and serve as our primary screen for potential drug candidates. The anti-hydroxylated lysine stain would further validate that the correction to collagen organization is due to a correction specific to collagen cross-linking. Importantly, this assay would be performed on fibroblasts cultured to recapitulate the more physiologically-relevant 3D environment of the skin tissue, rather than in the more artificial environment of the 2D monolayer, which can serve as the backup option.
To follow up on our drug candidates identified in the primary repurposing screen, we will further characterize the ECM of treated fibroblasts to ensure that healthy collagen maturation is taking place. We will assess cell health by using an Alamar Blue stain to quantify metabolic activity in the tissue (Rønnow et al., 2020). Electrophoretic mobility of collagen isolated from these tissues will serve as a secondary readout for restored collagen cross-linking through lysine hydroxylation. Finally, although there was no significant difference in tensile strength between control and PLOD1-deficient fibroblasts in 3D culture, we could opt to take these measurements in treated samples with an nano-indenter (Micha et al., 2019). This work can be done by one or more academic labs after Charles River Laboratories completes the primary drug screen and we have a list of validated hits in-hand.
Validation of drug candidates on additional cell lines of affected tissues
Given the involvement of multiple organ systems in PLOD1-kEDS, it is important to assess efficacy of a repurposed drug on more than one affected tissue. Vascular Smooth Muscle Cell (VSMC) cultures are validated models for studying the LH1 enzyme activity and are derived from key tissues that are affected in PLOD1-kEDS. We propose using the same image-based readouts to test drug candidates in this additional cellular context to assess collagen maturation in blood vessel tissue. Finding a drug that increases the stability of the extracellular matrix in blood vessels would be a boon given the arterial fragility and risk for rupture that is characteristic of the disorder.
FUNDING MODEL
XXXXX’s family has provided the seed money for Year One activities of the PLOD1-kEDS Cure Roadmap project. Data generated from Cure Roadmap activities will then serve as a basis for the fundraising necessary for subsequent drug development.
OPERATING STRUCTURE
Perlara Cure Guides will collaborate closely with XXXXX’s family. If at some point in the future it makes scientific and business sense, Perlara and XXXXX’s family will explore becoming co-development partners in order to create joint ventures to bring medicines to market.
BEYOND PLOD1-kEDS
The therapeutic modalities tested and subsequently developed to treat XXXXX’s PLOD1-kEDS have broader applicability. Each of the therapeutic tracks described can be applicable to other PLOD1-kEDS variants and further expanded to treat a larger subset of Ehlers-Danlos syndromes. For example, the high-throughput screening strategy used for drug repurposing can be used to identify therapies for Bruck Syndrome, Stickler syndrome-like connective tissue disorder (Ewans et al., 2019), and Spondylocheiro dysplastic EDS (Giunta et al., 2008). Furthermore, the knowledge base used in developing therapies that target vascular smooth muscle is directly applicable for treating the several EDS subtypes that affect the vasculature (Malfait, 2018).
Kyphoscoliotic Ehler-Danlos syndrome was the first subtype of connective tissue disorders for which the causative genetic mutation was identified (Pinnell et al., 1972), a discovery subsequently that paved the way for the comprehensive characterization and management of the Ehlers-Danlos syndromes. We believe that the treatment avenues proposed for treating XXXXX’s n-of-1 variant serve as the equivalent spark needed to advance development of therapeutics for a much broader range of connective tissue health. Furthermore, we see this research being critically informative for addressing the unmet needs of many suffering from the effects of aging and vascular, muscle, and bone disease.
ROADMAP UPDATES
Will be added as the project progresses.
REFERENCES
Askou AL, Jakobsen TS, Corydon TJ. Retinal gene therapy: an eye-opener of the 21st century. Gene Ther. 2021 May;28(5):209-216. doi: 10.1038/s41434-020-0168-2. Epub 2020 Jun 19. PMID: 32561864.
Bainbridge JW, Tan MH, Ali RR. Gene therapy progress and prospects: the eye. Gene Ther. 2006 Aug;13(16):1191-7. doi: 10.1038/sj.gt.3302812. Epub 2006 Jul 13. PMID: 16838031.
Bellen HJ, Levis RW, Liao G, He Y, Carlson JW, Tsang G, Evans-Holm M, Hiesinger PR, Schulze KL, Rubin GM, Hoskins RA, Spradling AC. The BDGP gene disruption project: single transposon insertions associated with 40% of Drosophila genes. Genetics. 2004 Jun;167(2):761-81. doi: 10.1534/genetics.104.026427. PMID: 15238527; PMCID: PMC1470905.
Chiba S, Lim KRQ, Sheri N, Anwar S, Erkut E, Shah MNA, Aslesh T, Woo S, Sheikh O, Maruyama R, Takano H, Kunitake K, Duddy W, Okuno Y, Aoki Y, Yokota T. eSkip-Finder: a machine learning-based web application and database to identify the optimal sequences of antisense oligonucleotides for exon skipping. Nucleic Acids Res. 2021 Jul 2;49(W1):W193-W198. doi: 10.1093/nar/gkab442. PMID: 34104972; PMCID: PMC8265194.
Ewans LJ, Colley A, Gaston-Massuet C, Gualtieri A, Cowley MJ, McCabe MJ, Anand D, Lachke SA, Scietti L, Forneris F, Zhu Y, Ying K, Walsh C, Kirk EP, Miller D, Giunta C, Sillence D, Dinger M, Buckley M, Roscioli T. Pathogenic variants in PLOD3 result in a Stickler syndrome-like connective tissue disorder with vascular complications. J Med Genet. 2019 Sep;56(9):629-638. doi: 10.1136/jmedgenet-2019-106019. Epub 2019 May 25. PMID: 31129566.
Ghali N, Sobey G, Burrows N. Ehlers-Danlos syndromes. BMJ. 2019 Sep 18;366:l4966. doi: 10.1136/bmj.l4966. PMID: 31533917.
Giunta C, Baumann M, Fauth C, Lindert U, Abdalla EM, Brady AF, Collins J, Dastgir J, Donkervoort S, Ghali N, Johnson DS, Kariminejad A, Koch J, Kraenzlin M, Lahiri N, Lozic B, Manzur AY, Morton JEV, Pilch J, Pollitt RC, Schreiber G, Shannon NL, Sobey G, Vandersteen A, van Dijk FS, Witsch-Baumgartner M, Zschocke J, Pope FM, Bönnemann CG, Rohrbach M. A cohort of 17 patients with kyphoscoliotic Ehlers-Danlos syndrome caused by biallelic mutations in FKBP14: expansion of the clinical and mutational spectrum and description of the natural history. Genet Med. 2018 Jan;20(1):42-54. doi: 10.1038/gim.2017.70. Epub 2017 Jun 15. PMID: 28617417; PMCID: PMC5763155.
Graur M, Ecovoiu AAL, Ratiu AC, Savu L, Gavrila L. CG6199LH2a lethal allele from Drosophila melanogaster is a candidate model for investigations on Ehlers-Danlos syndrome. Roumanian Biotechnological Letters. 2008. Oct;13(1):4066-4073.
Heikkinen J, Toppinen T, Yeowell H, Krieg T, Steinmann B, Kivirikko KI, Myllylä R. Duplication of seven exons in the lysyl hydroxylase gene is associated with longer forms of a repetitive sequence within the gene and is a common cause for the type VI variant of Ehlers-Danlos syndrome. Am J Hum Genet. 1997 Jan;60(1):48-56. PMID: 8981946; PMCID: PMC1712545.
Ishikawa Y, Bächinger HP. A molecular ensemble in the rER for procollagen maturation. Biochim Biophys Acta. 2013 Nov;1833(11):2479-91. doi: 10.1016/j.bbamcr.2013.04.008. Epub 2013 Apr 18. PMID: 23602968.
Ishikawa Y, Mizuno N, Holden P, Lim PJ, Gould DB, Rohrbach M, Giunta C, Bächinger HP. The novel missense mutation Met48Lys in FKBP22 changes its structure and functions. Sci Rep. 2020 Jan 16;10(1):497. doi: 10.1038/s41598-019-57374-y. PMID: 31949249; PMCID: PMC6965642.
Jarisch A, Giunta C, Zielen S, König R, Steinmann B. Sibs affected with both Ehlers-Danlos syndrome type IV and cystic fibrosis. Am J Med Genet. 1998 Aug 6;78(5):455-60.doi:10.1002/(sici)1096-8628(19980806)78:5<455::aid-ajmg11>3.0.co;2-e. PMID: 9714013.
Kakkis ED. Enzyme replacement therapy for the mucopolysaccharide storage disorders. Expert Opin Investig Drugs. 2002 May;11(5):675-85. doi: 10.1517/13543784.11.5.675. PMID: 11996648.
Krane SM, Pinnell SR, Erbe RW. Lysyl-protocollagen hydroxylase deficiency in fibroblasts from siblings with hydroxylysine-deficient collagen. Proc Natl Acad Sci U S A. 1972 Oct;69(10):2899-903. doi: 10.1073/pnas.69.10.2899. PMID: 4342967; PMCID: PMC389670.
Krol BJ, Murad S, Walker LC, Marshall MK, Clark WL, Pinnell SR, Yeowell HN. The expression of a functional, secreted human lysyl hydroxylase in a baculovirus system. J Invest Dermatol. 1996 Jan;106(1):11-6. doi: 10.1111/1523-1747.ep12326956. PMID: 8592059.
Li D, Mastaglia FL, Fletcher S, Wilton SD. Precision Medicine through Antisense Oligonucleotide-Mediated Exon Skipping. Trends Pharmacol Sci. 2018 Nov;39(11):982-994. doi: 10.1016/j.tips.2018.09.001. Epub 2018 Sep 30. PMID: 30282590.
Malfait F. Vascular aspects of the Ehlers-Danlos Syndromes. Matrix Biol. 2018 Oct;71-72:380-395. doi: 10.1016/j.matbio.2018.04.013. Epub 2018 Apr 27. PMID: 29709596.
Micha D, Pals G, Smit TH, Ghazanfari S. An in vitro model to evaluate the properties of matrices produced by fibroblasts from osteogenesis imperfecta and Ehlers-Danlos Syndrome patients. Biochem Biophys Res Commun. 2020 Jan 8;521(2):310-317. doi: 10.1016/j.bbrc.2019.09.081. Epub 2019 Oct 24. PMID: 31668813.
Newbrun E. Indices to measure gingival bleeding. J Periodontol. 1996 Jun;67(6):555-61. doi: 10.1902/jop.1996.67.6.555. PMID: 8794964.
Norman KR, Moerman DG. The let-268 locus of Caenorhabditis elegans encodes a procollagen lysyl hydroxylase that is essential for type IV collagen secretion. Dev Biol. 2000 Nov 15;227(2):690-705. doi: 10.1006/dbio.2000.9897. PMID: 11071784.
Parenti G, Andria G, Ballabio A. Lysosomal storage diseases: from pathophysiology to therapy. Annu Rev Med. 2015;66:471-86. doi: 10.1146/annurev-med-122313-085916. PMID: 25587658.
Pinnell SR, Krane SM, Kenzora JE, Glimcher MJ. A heritable disorder of connective tissue. Hydroxylysine-deficient collagen disease. N Engl J Med. 1972 May 11;286(19):1013-20. doi: 10.1056/NEJM197205112861901. PMID: 5016372.
Prater SN, Banugaria SG, DeArmey SM, Botha EG, Stege EM, Case LE, Jones HN, Phornphutkul C, Wang RY, Young SP, Kishnani PS. The emerging phenotype of long-term survivors with infantile Pompe disease. Genet Med. 2012 Sep;14(9):800-10. doi: 10.1038/gim.2012.44. Epub 2012 Apr 26. PMID: 22538254; PMCID: PMC3947503.
Qi Y, Xu R. Roles of PLODs in Collagen Synthesis and Cancer Progression. Front Cell Dev Biol. 2018 Jun 28;6:66. doi: 10.3389/fcell.2018.00066. PMID: 30003082; PMCID: PMC6031748.
Rønnow SR, Dabbagh RQ, Genovese F, Nanthakumar CB, Barrett VJ, Good RB, Brockbank S, Cruwys S, Jessen H, Sorensen GL, Karsdal MA, Leeming DJ, Sand JMB. Prolonged Scar-in-a-Jar: an in vitro screening tool for anti-fibrotic therapies using biomarkers of extracellular matrix synthesis. Respir Res. 2020 May 7;21(1):108. doi: 10.1186/s12931-020-01369-1. PMID: 32381012; PMCID: PMC7203825.
Ruiz-Botero F, Ramírez-Montaño D, Pachajoa H. FKBP14 kyphoscoliotic Ehlers-Danlos Syndrome in adolescent patient: the first Colombian report. Arch Argent Pediatr. 2019 Jun 1;117(3):e274-e278. English, Spanish. doi: 10.5546/aap.2019.eng.e274. PMID: 31063316.
Scietti L, Campioni M, Forneris F. SiMPLOD, a Structure-Integrated Database of Collagen Lysyl Hydroxylase (LH/PLOD) Enzyme Variants. J Bone Miner Res. 2019 Jul;34(7):1376-1382. doi: 10.1002/jbmr.3692. Epub 2019 Mar 12. PMID: 30721533.
Scietti L, Chiapparino A, De Giorgi F, Fumagalli M, Khoriauli L, Nergadze S, Basu S, Olieric V, Cucca L, Banushi B, Profumo A, Giulotto E, Gissen P, Forneris F. Molecular architecture of the multifunctional collagen lysyl hydroxylase and glycosyltransferase LH3. Nat Commun. 2018 Aug 8;9(1):3163. doi: 10.1038/s41467-018-05631-5. Erratum in: Nat Commun. 2018 Sep 20;9(1):3912. PMID: 30089812; PMCID: PMC6082870.
Uzawa K, Grzesik WJ, Nishiura T, Kuznetsov SA, Robey PG, Brenner DA, Yamauchi M. Differential expression of human lysyl hydroxylase genes, lysine hydroxylation, and cross-linking of type I collagen during osteoblastic differentiation in vitro. J Bone Miner Res. 1999 Aug;14(8):1272-80. doi: 10.1359/jbmr.1999.14.8.1272. PMID: 10457259.
Yeowell HN, Allen JD, Walker LC, Overstreet MA, Murad S, Thai SF. Deletion of cysteine 369 in lysyl hydroxylase 1 eliminates enzyme activity and causes Ehlers-Danlos syndrome type VI. Matrix Biol. 2000 Feb;19(1):37-46. doi: 10.1016/s0945-053x(99)00055-4. PMID: 10686424.
Yeowell HN, Steinmann B. PLOD1-Related Kyphoscoliotic Ehlers-Danlos Syndrome. 2000 Feb 2 [updated 2018 Oct 18]. In: Adam MP, Ardinger HH, Pagon RA, Wallace SE, Bean LJH, Mirzaa G, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2021. PMID: 20301635.
Yeowell HN, Walker LC. Mutations in the lysyl hydroxylase 1 gene that result in enzyme deficiency and the clinical phenotype of Ehlers-Danlos syndrome type VI. Mol Genet Metab. 2000 Sep-Oct;71(1-2):212-24. doi: 10.1006/mgme.2000.3076. PMID: 11001813.
Wahane A, Waghmode A, Kapphahn A, Dhuri K, Gupta A, Bahal R. Role of Lipid-Based and Polymer-Based Non-Viral Vectors in Nucleic Acid Delivery for Next-Generation Gene Therapy. Molecules. 2020 Jun 22;25(12):2866. doi: 10.3390/molecules25122866. PMID: 32580326; PMCID: PMC7356024.
Winkel LP, Kamphoven JH, van den Hout HJ, Severijnen LA, van Doorn PA, Reuser AJ, van der Ploeg AT. Morphological changes in muscle tissue of patients with infantile Pompe's disease receiving enzyme replacement therapy. Muscle Nerve. 2003 Jun;27(6):743-51. doi: 10.1002/mus.10381. PMID: 12766987.
Zhou T, Benda C, Dunzinger S, Huang Y, Ho JC, Yang J, Wang Y, Zhang Y, Zhuang Q, Li Y, Bao X, Tse HF, Grillari J, Grillari-Voglauer R, Pei D, Esteban MA. Generation of human induced pluripotent stem cells from urine samples. Nat Protoc. 2012 Dec;7(12):2080-9. doi: 10.1038/nprot.2012.115. Epub 2012 Nov 8. PMID: 23138349.